The same N stimuli would also be used as the inputs to the computational model. For each pair of stimuli, you can quantify the similarity of model’s response to the two stimuli (e.g., the correlation between the pattern of activation produced by the two stimuli). This gives you an N x N representational similarity matrix for the model.
Now we’ve transformed both the ERP data and the model results into N x N representational similarity matrices. The ERP data and the model originally had completely different units of measurement and data structures that were difficult to relate to each other, but now we have the same data format for both the ERPs and the model. This makes it simple to ask how well the similarity matrix for the ERP data matches the similarity matrix for the model. Specifically, we can just calculate the correlation between the two matrices (typically using a rank order approach so that we only assume a monotonic relationship, not a linear relationship).
Some Details
The data shown in Figure 1 used the Pearson r correlation coefficient to quantify the similarity between ERP scalp distributions. We have found that this is a good metric of similarity for ERPs, but other metrics can sometimes be advantagous. Note that many researchers prefer to quantify dissimilarity (distance) rather than similarity, but the principle is the same.
Each representational similarity matrix (RSM) captures the representational geometry of the system that produced the data (e.g., the human brain or the computational model). The lower and upper triangles of the RSM as described in this approach are mirror images of each other and are redundant. Similarly, cells along the diagonal index the similarity of each item to itself and are not considered in cross-RSM comparisons. We therefore use only the lower triangles of the RSMs. As illustrated in Figure 2a, the representational similarity between the ERP data and the computational model is simply the (rank order) correlation between the values in these two lower triangles.
When RSA is used with ERP data, representational similarity is typically calculated separately for each time point. That is, the scalp distribution is obtained at a given time point for each of the N stimuli, and the correlation between the scalp distributions for each pair of stimuli is computed at this time point. Thus, we have an N x N RSM at each time point for the ERP data. Each of these RSMs is then correlated with the RSM from the computational model. If the model has multiple layers, this process is conducted separately for each layer.
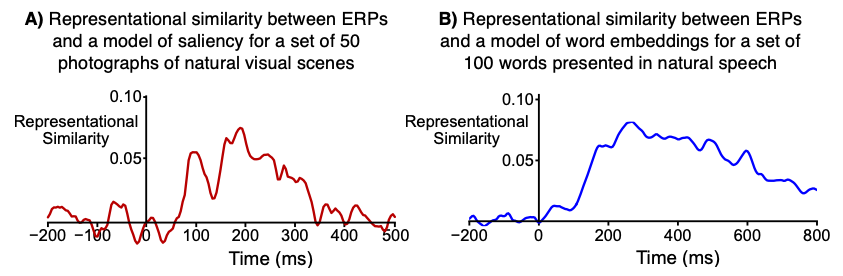
For example, the waveforms shown in Figure 1 show the (rank order) correlation between the ERP RSM at a given time point and the model RSM. That is, each time point in the waveform shows the correlation between the ERP RSM for that time point and the model RSM.
ERP scalp distributions can vary widely across people, so RSA is conducted separately for each participant. That is, we compute an ERP RSM for each participant (at each time point) and calculate the correlation between that RSM and the Model RSM. This gives us a separate ERP-Model correlation value for each participant at each time point. The waveforms shown in Figure 1 show the average of the single-participant correlations.
The correlation values in RSA studies of ERPs are typically quite low compared to the correlation values you might see in other contexts (e.g., the correlation between P3 latency and response time). For example, all of the correlation values in the waveforms shown in Figure 1 are less than 0.10. However, this is not usually a problem for the following reasons:
The low correlations are mainly a result of the noisiness of scalp ERP data when you compute a separate ERP for each of 50-100 stimuli, not a weak link between the brain and the model.
It is possible to calculate a “noise ceiling,” which represents the highest correlation between RSMs that could be expected given the noise in the data. The waveforms shown in Figure 1 reach a reasonably high value relative to the noise ceiling.
When the correlation between the ERP RSM and the model RSM is computed for a given participant, the number of data points contributing to the correlation is typically huge. For a 50 x 50 RSM (as in Figure 1A), there are 1225 cells in the lower triangle. 1225 values from the ERP RSM are being correlated with 1225 values from the model RSM. This leads to very robust correlation estimates.
Additional power is achieved from the fact that a separate correlation is computed for each participant.
In practice, the small correlation values obtained in ERP RSA studies are scientifically meaningful and can have substantial statistical power.
RSA is usually applied to averaged ERP waveforms, not single-trial data. For example, we used averages of 32 trials per image in the experiment shown in Figure 1A. The data shown in Figure 1B are from averages of at least 10 trials per word. Single-trial analyses are possible but are much noisier. For example, we conducted single-trial analyses of the words and found statistically significant but much weaker representational similarity.
Other Types of Data
As illustrated in Figure 2A, RSA can also be used to link ERPs to other types of data, including behavioral data and fMRI data.
The behavioral example in Figure 2A involves eye tracking. If the eyes are tracked while participants view scenes, a fixation density map can be constructed showing the likelihood that each location was fixated for each scene. An RSM for the eye-tracking data could be constructed to indicate the similarity between fixation density maps for each pair of scenes. This RSM could then be correlated with the ERP RSM at each time point. Or the fixation density RSMs could be correlated with the RSM for a computational model (as in a recent study in which we examined the relationship between infant eye movement patterns and a convolutional neural network model of the visual system; Kiat et al., 2022).
Other types of behavioral data could also be used. For example, if participants made a button-press response to each stimulus, one could use the mean response times for each stimulus to construct an RSM. The similarity value for a given cell would be the difference in mean RT between two different stimuli.
RSA can also be used to link ERP data to fMRI data, a process called data fusion (see, e.g., Mohsenzadeh et al., 2019). The data fusion process makes it possible to combine the spatial resolution of fMRI with the temporal resolution of ERPs. It can yield a millisecond-by-millisecond estimate of activity corresponding to a given brain region, and it can also yield a voxel-by-voxel map of the activity corresponding to a given time point. More details are provided in our YouTube video on RSA.